1. De Bruijn graphs and kmer sets
- GGCAT: Extremely-fast construction and querying of compacted and colored de Bruijn graphs
- matchtigs & eulertigs: Minimum plain-text representation of kmer sets / spectrum preserving string sets
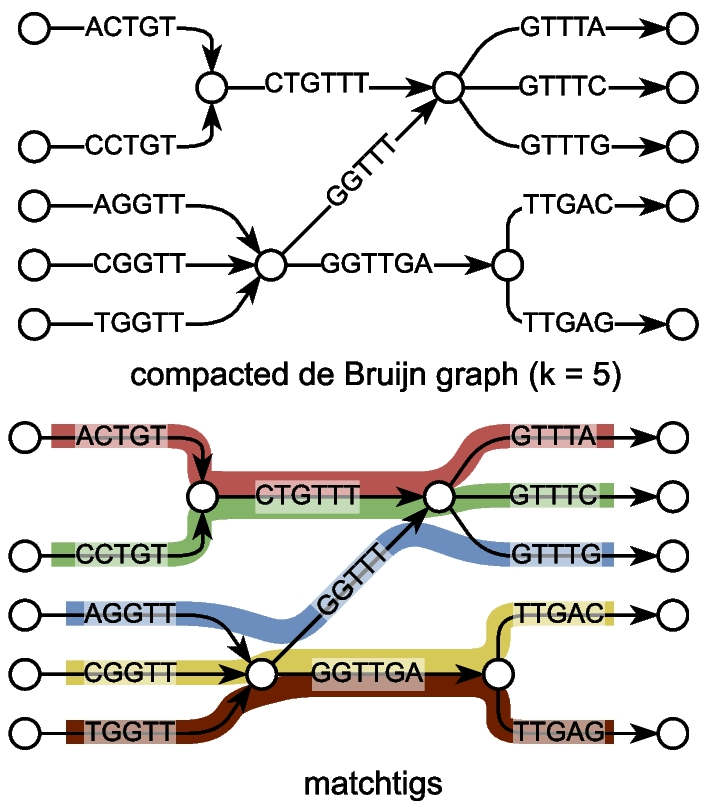
1. De Bruijn graphs and kmer sets
|

|
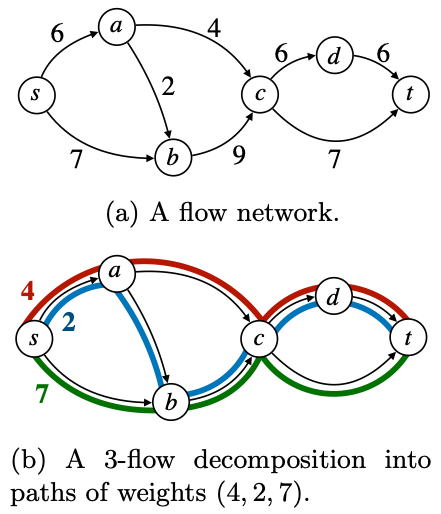
2. Fast solvers for flow decomposition problems via Integer Linear Programming
|
|
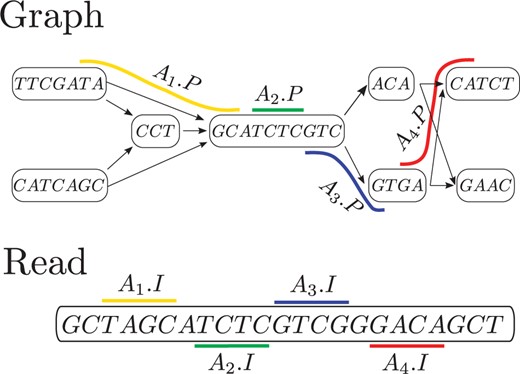
3. Alignment to pangenomes
|
|
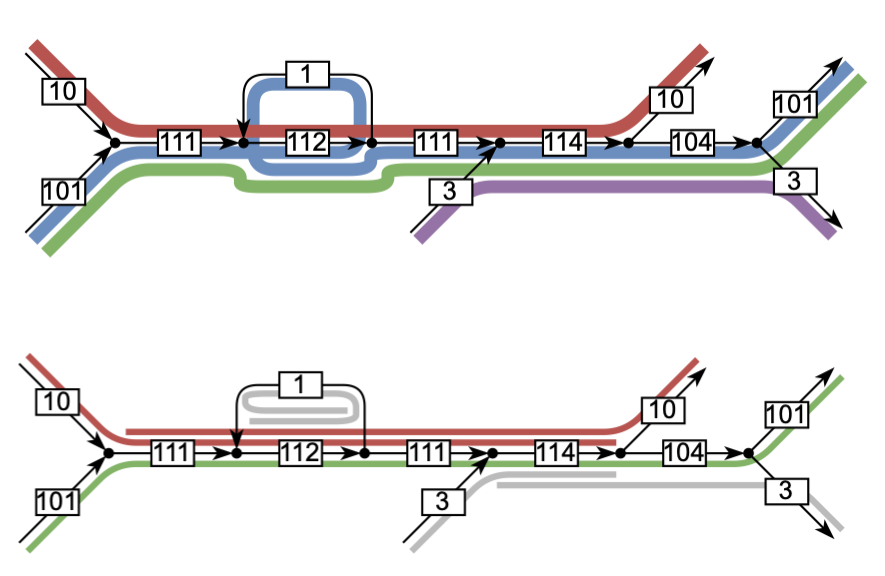
4. Super-unitigs for genome assembly
|
|
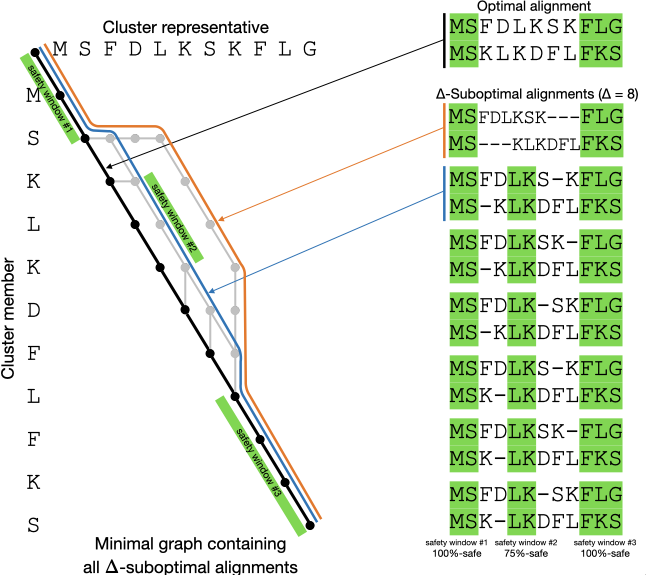
5. Protein and RNA alignment
|
|